Quantitative Polymerase Chain Reaction (qPCR)
Quantitative polymerase chain reaction (qPCR) quantifies the abundance of specific microorganisms and functional genes capable of degrading a particular contaminant at a given site. Thus, qPCR-related analytical approaches can be used to evaluate contamination clean-up efforts. Here, we outline the basics, list the main approaches, discuss their advantages / disadvantages, consider when to use them, provide guidance on sampling, and review several case studies applied to sites with chlorinated solvents, petroleum hydrocarbons, and perchlorate.
Related Article(s):
- Compound Specific Isotope Analysis (CSIA)
- Metagenomics
- Molecular Biological Tools - MBTs
- Stable Isotope Probing (SIP)
Contributor(s): Dora Ogles-Taggart and Dr. Brett Baldwin
Key Rescource(s):
- Next Generation qPCR: High Throughput, Highly Parallel qPCR Arrays (QuantArrays) for Comprehensive Site Assessment[1]
- Quantitative Real-Time PCR (qPCR) Detection Chemistries Affect Enumeration of the Dehalococcoides 16S rRNA Gene in Groundwater[2]
Introduction
Quantitative polymerase chain reaction (qPCR) measures the concentration (abundance) of specific microorganisms or functional genes that are capable of degrading a particular contaminant. High concentrations of contaminant degrading microorganisms (e.g., Dehalococcoides) or specific functional genes (e.g., vinyl chloride reductase) is a direct indicator of in situ biodegradation potential. Conversely, low concentrations of contaminant degrading microorganisms suggest that amendments may be needed to promote biodegradation. Therefore, results from qPCR analysis can be an important line of evidence when evaluating monitored natural attenuation (MNA) and bioremediation strategies.
- qPCR: A DNA-based technique used to detect and quantify specific microorganisms or functional genes that can biodegrade contaminants of concern.
- Reverse transcriptase quantitative polymerase chain reaction (RT-qPCR): A similar technique that quantifies biodegradation activity. When a microorganism is biodegrading a target contaminant, genes for the enzymes needed for biodegradation are being expressed – DNA is being transcribed into the corresponding mRNA sequence. By measuring RNA, RT-qPCR quantifies the expression of the target functional genes. Results therefore indicate biodegradation activity.
- QuantArray: A qPCR or RT-qPCR tool that can simultaneously quantify a broad suite of target genes or their transcription (indicating biological activity) in a single analysis to provide a more comprehensive evaluation of contaminant biodegradation[1].
Advantages
qPCR is a technique to quantify microorganisms of interest without having to grow/cultivate them in the laboratory using traditional approaches like plate counts. Less than 1% of bacteria have been cultivated in a laboratory[3]; plate count results are therefore biased and may not be suitable to enumerate organisms. Other advantages of qPCR are as follows:
- Accurate: qPCR provides an accurate quantification of contaminant-degrading microorganisms and relevant functional genes based on the analysis of DNA extracted directly from a soil or groundwater sample. While not an Environmental Protection Agency (EPA) method, standard protocols have been developed for sample collection, preservation, and processing to ensure accurate quantification[2][4][5].
- Sensitive: The detection limit is typically 100 cells or gene copies per sample. This is orders of magnitude lower than concentrations of contaminant degraders generally needed for effective bioremediation. For example, a threshold of 104 Dehalococcoides cells/mL (107 cells per 1 L sample) is recommended for generally effective rates of reductive dechlorination at chlorinated ethene sites[2].
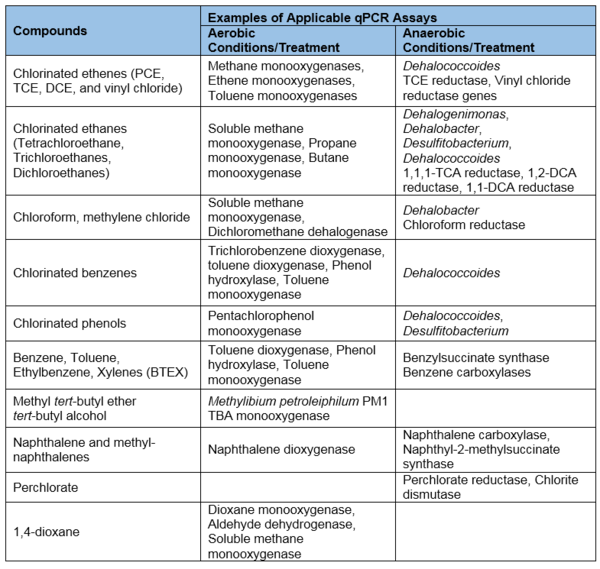
- Broadly applicable: qPCR can be used for a variety of common soil and groundwater contaminants, including chlorinated solvents, petroleum hydrocarbons, perchlorate, 1,4-dioxane, and more (Table 1). This tabulation reflects the results of over 15 years of research to identify the groups of microorganisms and functional genes responsible for contaminant biodegradation.
- Activity indicator: RT-qPCR quantifies gene expression and is therefore used to evaluate activity.
Limitations
- Unknown microorganisms and biodegradation pathways: qPCR assays can only be developed if the contaminant biodegradation pathway or target microorganism is known (Table 1). As research progresses, our understanding of contaminant biodegradation will improve and qPCR assays can likewise be expanded to apply to new contaminants, microorganisms, pathways, and gene sequences.
- PCR inhibition: Although uncommon, high concentrations of humic acids and some heavy metals can inhibit qPCR. In practice, PCR inhibition would be identified by reviewing quality control parameters and can often be overcome through standard operating procedures such as performing dilutions[3].
When to Use qPCR
Simply put, high concentrations of contaminant degrading microorganisms and functional genes suggests that biodegradation is more likely whereas low concentrations of contaminant degraders suggest biodegradation is limited under the existing conditions. Therefore, submitting samples for qPCR analysis should be strongly considered during remedy selection and performance monitoring to aid in answering the following types of questions:
- Site Assessment/Remedy Selection:
- MNA: Is MNA feasible? Are contaminant degrading microorganisms present at high concentrations under existing site conditions?
- Biostimulation: Should an amendment such as an electron donor or acceptor be added to stimulate growth of contaminant degrading microorganisms?
- Bioaugmentation: Should a commercial biological culture be added to ensure that contaminant-degrading microorganisms are present at high concentrations?
- Performance Monitoring:
- MNA: Are contaminant-degrading populations maintained over time?
- Biostimulation: Was amendment addition effective? Did it promote growth of contaminant-degrading microorganisms? Are contaminant degrader populations maintained over time? Is another amendment addition warranted?
- Bioaugmentation: Did the bioaugmentation culture survive in situ? Are contaminant degrader populations maintained over time?
Use at Chlorinated Solvent Sites
Anaerobic bioremediation by biostimulation or bioaugmentation is a common treatment strategy at sites impacted by chlorinated solvents such as tetrachloroethene (PCE) and trichloroethene (TCE). To date, Dehalococcoides remains the only known bacterial group capable of complete reductive dechlorination of PCE and TCE through cis- dichloroethene (cis-DCE) and vinyl chloride to ethene. Therefore, qPCR quantification of Dehalococcoides and vinyl chloride reductase genes has become a routine component of assessment, remedy selection, and performance monitoring at sites impacted by chlorinated solvents. In fact, a threshold concentration of 104 Dehalococcoides cells/mL is proposed for generally effective rates of reductive dechlorination at sites impacted by chlorinated ethenes[6].
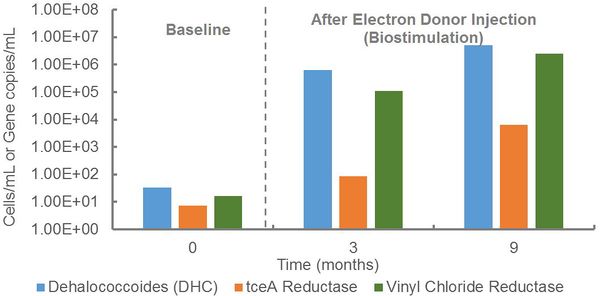
At an industrial site, qPCR was performed to evaluate the potential for anaerobic biodegradation of a mixture of chlorinated solvents (Fig. 1). During site assessment, Dehalococcoides and vinyl chloride reductase genes were detected at low concentrations, indicating the presence of organohalide-respiring bacteria capable of complete reductive dechlorination of TCE to ethene. In response to electron donor injection, Dehalococcoides populations and vinyl chloride reductase gene copies increased by more than four orders of magnitude to concentrations exceeding the 104 cells/mL recommended for generally effective rates of reductive dechlorination[6]. Thus, electron donor stimulated growth of this key group of organohalide-respiring bacteria and promoted anaerobic biodegradation.
Under oxic conditions, several different types of bacteria such as methanotrophs and etheneotrophs can cometabolically degrade TCE and lesser chlorinated ethenes to non-toxic products. In general, cometabolism is a result of monooxygenase enzymes that oxidize a primary substrate (e.g., methane) to support growth of the microorganism but have broad substrate specificity that permits co-oxidation of chlorinated ethenes. qPCR can also quantify concentrations of methane monooxygenase genes and ethene monooxygenase genes to evaluate the potential for cometabolic biodegradation of TCE[7].
Use at Petroleum Hydrocarbon Sites
Aerobic biodegradation of petroleum hydrocarbons including benzene, toluene, ethylbenzene, and xylenes (BTEX), polycyclic aromatic hydrocarbons (PAHs), and alkanes is typically initiated by oxygenase enzymes (see[8][9] for review). Many qPCR assays targeting these oxygenase genes have been developed[10] to quantify aerobic BTEX and PAH degraders and employed to evaluate the potential for aerobic biodegradation during MNA or enhanced aerobic bioremediation[11][12]. In addition, RT-qPCR assays have been developed to quantify expression of oxygenase genes to more directly assess aerobic biodegradation activity[13][14].
Pathways and the corresponding genes have also been identified for anaerobic BTEX and naphthalene biodegradation. For toluene, ethylbenzene, and xylenes (alkyl substituted aromatics), anaerobic biodegradation is initiated by a benzylsuccinate synthase enzyme. qPCR assays have been developed to quantify the corresponding benzylsuccinate synthase (bssA) gene to evaluate anaerobic aromatic hydrocarbon biodegradation[15][16][17]. More recently, carboxylase genes have been implicated in initiating anaerobic biodegradation of benzene and naphthalene[18][19].
Use at Perchlorate Sites
Perchlorate is not readily absorbed, volatilized, or abiotically degraded making biodegradation the most important attenuation mechanism. Under anoxic conditions, perchlorate can serve as an electron acceptor and is sequentially biodegraded to chlorite and then to chloride and oxygen[20][21]. qPCR assays targeting perchlorate reductase and chlorite dismutase genes can be an important line of evidence to assess biodegradation of perchlorate[22].
Sampling Locations
As with any site assessment tool, selection of sampling locations is a critical step. Here are a few guidelines for selecting sampling locations to ensure that the results aid in meaningful site management decisions:
- Background samples: Comparison of qPCR results for non-impacted background samples results with impacted samples provides stronger evidence, particularly at petroleum hydrocarbon sites. Higher concentrations of contaminant-degrading microorganisms or functional genes (e.g., bssA) in samples from impacted areas relative to the background samples indicate selection and growth of contaminant degraders within the contaminant plume.
- Baseline samples: As with background samples, qPCR results from baseline samples collected and analyzed prior to treatment serve as the basis for comparison to evaluate growth of contaminant-degrading bacteria in response to the selected remediation strategy.
- Impacted samples: Sampling locations that represent distinct zones within the contaminant plume based on contaminant concentrations and geochemical conditions.
Sample Collection, Preservation, and Shipping
Sampling procedures for qPCR analysis are straightforward and are readily integrated into existing monitoring programs. Almost any type of sample matrix (soil, sediment, groundwater, on site filters) can be analyzed with qPCR. All samples should be shipped to the laboratory on ice (4°C) using an overnight carrier to minimize the potential for changes of the biomarkers of interest.
Groundwater samples (typically 1 L) can be shipped directly to the laboratory or filtered in the field. For on-site filtration, groundwater is pumped through a Sterivex® or Bio-Flo® filter using standard low flow sampling techniques. The volume of groundwater filtered is recorded so that the target gene abundance per liter can be calculated. The groundwater may then be discarded appropriately. For RNA analysis (RT-qPCR), a preservative is injected into the filter cartridge immediately after sample collection.
Summary
qPCR can accurately quantify key microorganisms and functional genes responsible for biodegrading common soil and groundwater contaminants, including chlorinated solvents, petroleum hydrocarbons and more. Submitting samples for qPCR analysis should be strongly considered during remedy selection and performance monitoring. High concentrations of contaminant degrading microorganisms and functional genes particularly when compared to background or baseline populations suggests that biodegradation is more likely. Conversely, low concentrations of contaminant degraders suggest biodegradation is limited under the existing conditions and amendments may be needed for successful bioremediation. Over the past 15 years, use of qPCR for assessing biodegradation during site assessment, remedy selection, and performance monitoring has increased dramatically. Routine qPCR quantification of Dehalococcoides has become commonplace at sites impacted by chlorinated solvents.
References
- ^ 1.0 1.1 Ogles, D.M., Biernacki, A., Baldwin, B.R., Ritalahti, K.M., Loeffler, F.E. 2014. Next Generation qPCR: High Throughput, Highly Parallel qPCR Arrays (QuantArrays) for Comprehensive Site Assessment. Report pdf
- ^ 2.0 2.1 2.2 Hatt, J.K. and Löffler, F.E., 2012. Quantitative real-time PCR (qPCR) detection chemistries affect enumeration of the Dehalococcoides 16S rRNA gene in groundwater. Journal of Microbiological Methods, 88(2), 263-270. doi: 10.1016/4.mimet.2011.12.005
- ^ 3.0 3.1 Amann, R.I., Ludwig, W. and Schleifer, K.H., 1995. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiological Reviews, 59(1), 143-169. Journal Article
- ^ Hatt, J.K., Ritalahti, K.M., Ogles, D.M., Lebrón, C.A., Löffler, F.E., 2013. Design and application of an internal amplification control to improve Dehalococcoides mccartyi 16S rRNA gene enumeration by qPCR. Environmental Science & Technology, 47(19), 11131-11138. doi: 10.1021/es4019817
- ^ Lebrón, C.A., Dennis, P., Acheson, C., Barros, N., Major, D., Petrovskis, E., Loffler, F.E., Ritalahti, K.M., Yeager, C.M., Edwards, E.A., Hatt, J.K., Ogles, D.M., 2014. Standardized procedures for use of nucleic acid-based tools - Recommendations for groundwater sampling and analysis using qPCR. ER-1561. Strategic Environmental Research Development Program, Arlington, VA. ER-1561
- ^ 6.0 6.1 Lu, X., Wilson, J.T., Kampbell, D.H., 2006. Relationship between Dehalococcoides DNA in ground water and rates of reductive dechlorination at field scale. Water Research, 40(16), 3131-3140. doi:10.1016/j.watres.2006.05.030
- ^ Mattes, T.E., Jin, Y.O., Dobson, M., Lee, M.C., Schmidt, S., Fogel, S., Findlay, M., Smoler, D., 2013. Quantifying the Presence and Activity of Aerobic, Vinyl Chloride-Degrading Microorganisms in Dilute Groundwater Plumes by Using Real-Time PCR. Project ER-1683. Strategic Environmental Research and Development Program, Arlington, VA. ER-1683
- ^ Cao, B., Nagarajan, K., Loh, K.C., 2009. Biodegradation of aromatic compounds: current status and opportunities for biomolecular approaches. Applied Microbiology and Biotechnology, 85(2), 207-228. doi:10.1007/s00253-009-2192-4
- ^ Wentzel, A., Ellingsen, T.E., Kotlar, H.K., Zotchev, S.B., Throne-Holst, M., 2007. Bacterial metabolism of long-chain n-alkanes. Applied microbiology and biotechnology, 76(6), 1209-1221. doi:10.1007/s00253-007-1119-1
- ^ Iwai, S., Johnson, T.A., Chai, B., Hashsham, S.A., Tiedje, J.M., 2011. Comparison of the specificities and efficacies of primers for aromatic dioxygenase gene analysis of environmental samples. Applied and Environmental Microbiology, 77(11), 3551-3557. doi: 10.1128/AEM.00331-11
- ^ Baldwin, B.R., Nakatsu, C.H., Nies, L., 2008. Enumeration of aromatic oxygenase genes to evaluate monitored natural attenuation at gasoline-contaminated sites. Water Research, 42(3), 723-731. doi:10.1016/j.watres.2007.07.052
- ^ DeBruyn, J.M., Chewning, C.S., Sayler, G.S., 2007. Comparative quantitative prevalence of Mycobacteria and functionally abundant nidA, nahAc, and nagAc dioxygenase genes in coal tar contaminated sediments. Environmental Science & Technology, 41(15), 5426-5432. doi:10.1021/es070406c
- ^ Baldwin, B.R., Biernacki, A., Blair, J., Purchase, M.P., Baker, J.M., Sublette, K., Davis, G., Ogles, D., 2010. Monitoring gene expression to evaluate oxygen infusion at a gasoline-contaminated site. Environmental Science & Technology, 44(17), 6829-6834. doi:10.1021/es101356t
- ^ Key, K.C., Sublette, K.L., Johannes, T.W., Ogles, D., Baldwin, B., Biernacki, A., 2014. Assessing BTEX Biodegradation Potential at a Refinery Using Molecular Biological Tools. Groundwater Monitoring & Remediation, 34(1), 35-48. doi:10.1111/gwmr.12037
- ^ Beller, H.R., Kane, S.R., Legler, T.C., Alvarez, P.J., 2002. A real-time polymerase chain reaction method for monitoring anaerobic, hydrocarbon-degrading bacteria based on a catabolic gene. Environmental Science & Technology, 36(18), 3977-3984. doi:10.1021/es025556w
- ^ Sublette, K., Peacock, A., White, D., Davis, G., Ogles, D., Cook, D., Kolhatkar, R., Beckmann, D., Yang, X., 2006. Monitoring Subsurface Microbial Ecology in a Sulfate‐Amended, Gasoline‐Contaminated Aquifer.Groundwater Monitoring & Remediation, 26(2), 70-78. doi: 10.1111/j.1745-6592.2006.00072.x
- ^ Winderl, C., Schaefer, S. and Lueders, T., 2007. Detection of anaerobic toluene and hydrocarbon degraders in contaminated aquifers using benzylsuccinate synthase (bssA) genes as a functional marker. Environmental Microbiology, 9(4), 1035-1046. doi:10.1111/j.1462-2920.2006.01230.x
- ^ Abu Laban, N., Selesi, D., Rattei, T., Tischler, P., Meckenstock, R.U., 2010. Identification of enzymes involved in anaerobic benzene degradation by a strictly anaerobic iron‐reducing enrichment culture. Environmental Microbiology, 12(10), 2783-2796. doi:10.1111/j.1462-2920.2010.02248.x
- ^ Bergmann, F.D., Selesi, D., Meckenstock, R.U., 2011. Identification of new enzymes potentially involved in anaerobic naphthalene degradation by the sulfate-reducing enrichment culture N47. Archives of Microbiology, 193(4), 241-250. doi:10.1007/s00203-010-0667-4
- ^ Kengen, S.W., Rikken, G.B., Hagen, W.R., Van Ginkel, C.G., Stams, A.J., 1999. Purification and characterization of (per)chlorate reductase from the chlorate-respiring strain GR-1. Journal of Bacteriology, 181(21), 6706-6711. Article
- ^ Van Ginkel, C.G., Rikken, G.B., Kroon, A.G.M., Kengen, S.W.M., 1996. Purification and characterization of chlorite dismutase: a novel oxygen-generating enzyme. Archives of Microbiology, 166(5), 321-326. doi:10.1007/s002030050390
- ^ Lieberman, M.T., Borden, R.C., 2008. Natural attenuation of perchlorate in groundwater: Processes, tools and monitoring techniques. Project ER-0428. Strategic Environmental Research and Development Program, Arlington, VA. ER-200428